Flowchart for the Analysis of Multi-Wavelength Sedimentation Velocity Data¶
Step 1: Import Experimental data into UltraScan-III OpenAUC format¶
Note
Required for data from XLA or XLI, this step is performed automatically in the Optima AUC. Sedimentation velocity data should not be measured in absorbance mode, use intensity mode instead.
Import the experimental data: Utilities: Import Experimental Data
Confirm Investigator setting and local/database selection
Import Experimental Data from local disk
Edit Run Information, Select Lab/Rotor/Calibration
Enter a Label (verbose description for the run)
Select the corresponding project by clicking on the Project button. If there is no project, create a new project.
Confirm experiment type and optical detection system
Enter any comments, if applicable
Select instrument, rotor, rotor calibration and operator
- Click on Accept.
Edit the Description field if necessary
Navigate to the first channel and select the centerpiece type.
- Select the proper solution - Make sure that the solution contains at least one analyte and a buffer
If you have more than one triple, you can click on Apply to All but verify centerpiece and solution for each triple first. Also, check the Description field again to make sure the appropriate information is saved.
If data were collected in intensity mode, you will need to Define Reference Scans by selecting a short region from the air-to-air interface portion of the data.
For equilibrium data from 6-channel centerpieces you should separate each channel with the Define Subsets/Process Subsets functions.
Failed triples or empty triples can be excluded from the run by clicking on Drop Selected Triples.
When everything has been set you can Save the scans to database or disk.
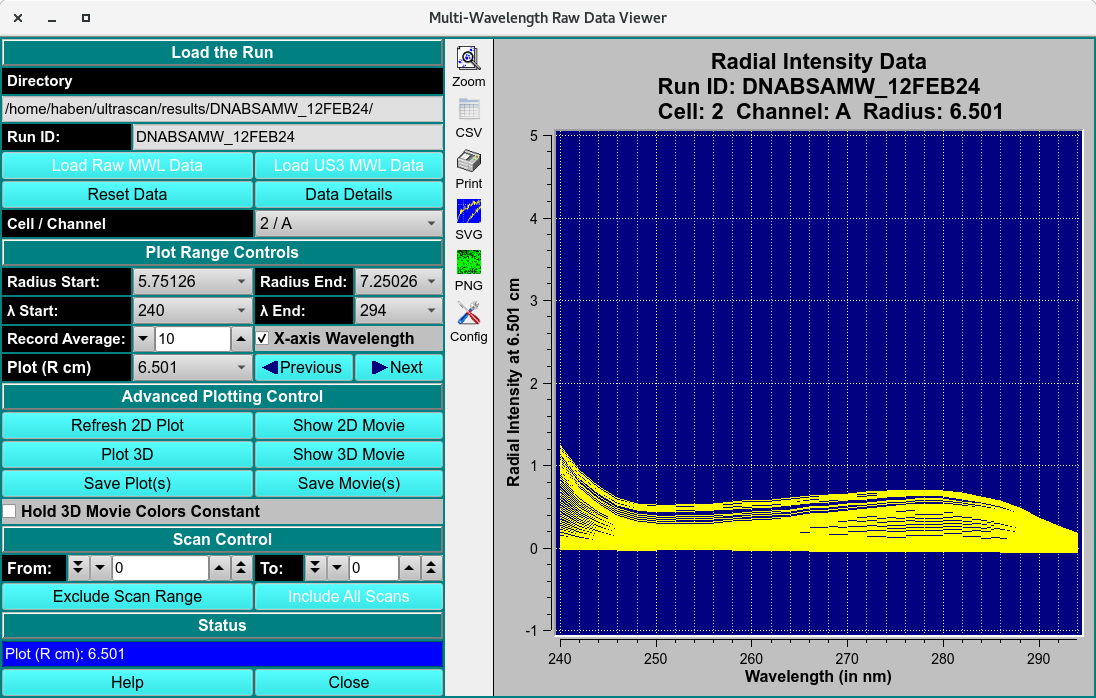
Alternate Record and X-axis
Step 2: Edit experimental data¶
Edit the data into Edit Data: Edit Data.
Load data from the database or a local data directory that contains the UltraScan 3 data files previously converted from the Beckman raw data.
Select the Cell / Channel / Wavelength triple to be edited.
Specify the meniscus of the data by holding down the Control key and using the left mouse button. The meniscus value may be manually adjusted with the keyboard.
If the data were collected with the interference detector, specify the left and right edges of the air gap area of the data.
Specify the left and right edges of the data to be analyzed.
Note
Please note: Do not pick the left data edge too close to the meniscus. During meniscus fitting, the evaluated meniscus positions may reach inside of the data range and violate the boundary conditions of the finite element solution. This will cause the meniscus fit to fail.
Specify the location of the scan plateau. This is the radial position where most scans have a stable plateau, but the selected position should not reach into the back-diffusion region. The most appropriate point tends to be close to the right edge of the data range, but not so far to the right that it extends into the region where the concentration of the later scans curves upward at the bottom of the cell due to back-diffusion.
Make any other optional adjustments to the data that are necessary and save the edit profile. When saving, a pop-up message is presented asking for an edit ID. The default for this ID is the current date and time in the form of YYMMddhhmm (Year / Month / Day / Hour / Minute), but this default can be supplemented with a suffix of your own choice.

Alternate Record and X-axis
Step 3: Inspect raw data¶
view raw mw data - mwlr_viewer.rst
Step 4: Perform a 2DSA refinement following the 2DSA Flowchart¶
Step 5: Synchronize each 2DSA-it models to a consistent time-grid¶
MWL - Simulations.rst
Step 6: Create basis spectra for each species¶
Step 7: Deconvolute MW data using species basis spectra¶
Step 8: Perform a 2DSA with refinement only¶
Step 9: Perform van Holde - Weischet analysis - (recommended)¶
Open Velocity: Enhanced van Holde - Weischet.
Load the desired experiment, applying the noise files from Step 6 (the latest model).
Check Plateaus from 2DSA and Use Enhanced vHW.
Adjust Beck Diffusion Tolerance, Divisions, Data Smoothing, % of Boundary, and Boundary Position to desired values.
If appropriate, delete early scans to improve resolution and reduce noise. Only keep scans and boundary portions that contribute to well correlated line fits in the linear extrapolations.
Select groups, if appropriate, to generate weight averaged s-values for discrete species.
Display Distribution Plot and histogram.
Save Data and distributions.
Note
Refer to the van Holde - Weischet manual page for additional details.
Step 10: Overlay combined distributions - (recommended)¶
All van Holde - Weischet distributions and finite element models can be combined into a single plot for easy comparison.
Use Velocity: Combine Distribution Plots (vHW) for van Holde - Weischet plots.
Use Velocity: Combine Discrete Distributions for all finite element models (2DSA, GA, Monte Carlo).
Use Velocity: Combine Integral Distributions for all finite element models (2DSA, GA, Monte Carlo).
Reference¶
2DSA
Brookes E, Cao W, Demeler B. A two-dimensional spectrum analysis for sedimentation velocity experiments of mixtures with heterogeneity in molecular weight and shape. Eur Biophys J. (2010) 39(3):405-14.
Kim H, Brookes E, Cao W, Demeler B. Two-dimensional grid optimization for sedimentation velocity analysis in the analytical ultracentrifuge. Eur Biophys J. 2018 Oct;47(7):837-844. doi: 10.1007/s00249-018-1309-z. PMID: 29777290
Experimental Design
Demeler, B. (2024). Methods for the design and analysis of analytical ultracentrifugation experiments. Current Protocols, 4, e974. doi: 10.1002/cpz1.974
PCSA
Gorbet G., T. Devlin, B. Hernandez Uribe, A. K. Demeler, Z. Lindsey, S. Ganji, S. Breton, L. Weise-Cross, E.M. Lafer, E.H. Brookes, B. Demeler. A parametrically constrained optimization method for fitting sedimentation velocity experiments. Biophys. J. (2014) vol 106, 1741-50.
Genetic Algorithm
Brookes E, Demeler B. Parsimonious Regularization using Genetic Algorithms Applied to the Analysis of Analytical Ultracentrifugation Experiments. GECCO ‘07: Proceedings of the 9th annual conference on Genetic and evolutionary computation, London, July 7-11, 2007, 361–368, https://doi.org/10.1145/1276958.1277035, ACM 978-1-59593-697-4/07/0007
Monte Carlo
Demeler B and E. Brookes. Monte Carlo analysis of sedimentation experiments. Colloid Polym Sci (2008) 286(2) 129-137
Adaptive Space-Time Finite Element Solution
Cao, W and Demeler B. Modeling Analytical Ultracentrifugation Experiments with an Adaptive Space-Time Finite Element Solution for Multi-Component Reacting Systems. Biophys. J. (2008) 95(1):54-65