Self-association Simulator¶
This module is used to simulate binding isotherms for reversibly self-associating systems with up to three species (monomer, N-mer, M-mer). The program is started from the Simulation menu (Self-Association Equilibrium):
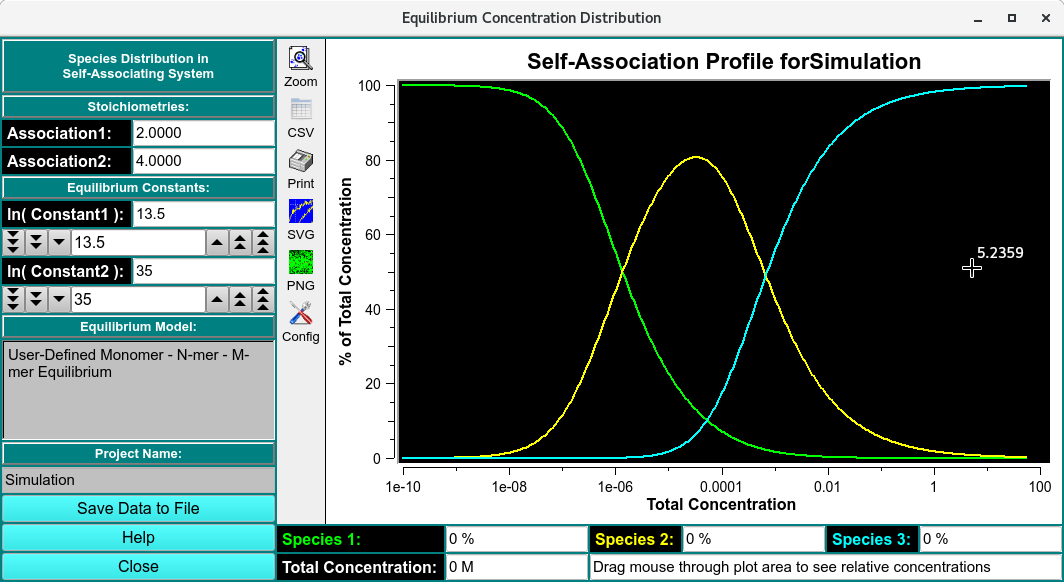
Self-Association Simulation Profile
Self-assoc. Simulator Functions:¶
Association (1,2): |
Enter an integral value to specify the stoichiometry of the association (e.g., 2, 3, 4…) |
Equilibrium constants: |
Enter the natural log of the association constants for each reaction. |
Save Data to File: |
Export the currently displayed association profile to a csv file. |
Self-Association Profile for Simulation: |
Binding isotherms for Monomer, N-mer and M-mer. |
Species 1/2/3: |
Percentage of species’ concentration with respect to the total concentration, requires mouse click on the canvas to be activated. |
Total Concentration: |
Total concentration of the reacting molecule. |
Help |
Display this and other documentation. |
Close |
Close all windows and exit. |